massing2022quantification
Quantification of metabolic niche occupancy dynamics in a Baltic Sea bacterial community
Jana C. Massing, Ashkaan K. Fahimipour, Carina Bunse, Jarone Pinhassi and Thilo Gross
mSystems 8, 28-23, 2023
Progress in molecular methods has enabled the monitoring of bacterial populations in time. Nevertheless, understanding community dynamics and its links with ecosystem functioning remains challenging due to the tremendous diversity of microorganisms. Conceptual frameworks that make sense of time-series of taxonomically-rich bacterial communities, regarding their potential ecological function, are needed. A key concept for organizing ecological functions is the niche, the set of strategies that enable a population to persist and define its impacts on the surroundings. Here we present a framework based on manifold learning, to organize genomic information into potentially occupied bacterial metabolic niches over time. We apply the method to re-construct the dynamics of putatively occupied metabolic niches using a long-term bacterial time-series from the Baltic Sea, the Linnaeus Microbial Observatory (LMO). The results reveal a relatively low-dimensional space of occupied metabolic niches comprising groups of taxa with similar functional capabilities. Time patterns of occupied niches were strongly driven by seasonality. Some metabolic niches were dominated by one bacterial taxon whereas others were occupied by multiple taxa, and this depended on season. These results illustrate the power of manifold learning approaches to advance our understanding of the links between community composition and functioning in microbial systems.
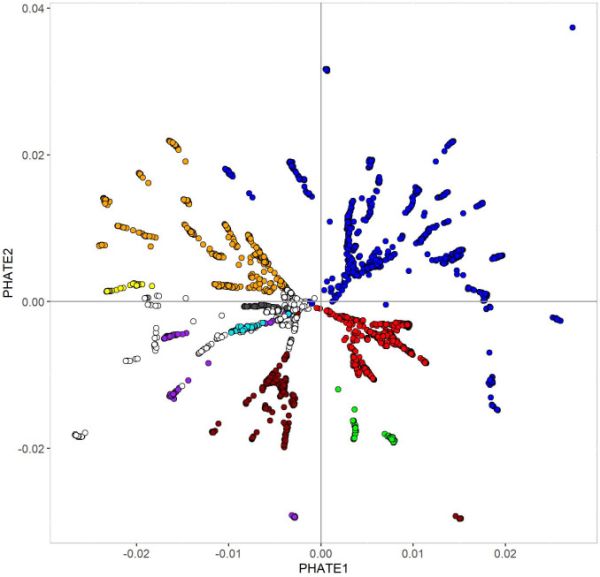
Figure 1: In this phate plot species (dots) are arranged according to their genetic similarity.